

correlation is an easystats
package focused on correlation analysis. It’s lightweight, easy to use,
and allows for the computation of many different kinds of correlations,
such as partial correlations, Bayesian
correlations, multilevel correlations,
polychoric correlations, biweight,
percentage bend or Sheperd’s Pi
correlations (types of robust correlation), distance
correlation (a type of non-linear correlation) and more, also allowing
for combinations between them (for instance, Bayesian partial
multilevel correlation).
You can cite the package as follows:
Makowski, D., Ben-Shachar, M. S., Patil, I., & Lüdecke, D. (2020). Methods and algorithms for correlation analysis in R. Journal of Open Source Software, 5(51), 2306. https://doi.org/10.21105/joss.02306
Makowski, D., Wiernik, B. M., Patil, I., Lüdecke, D., & Ben-Shachar, M. S. (2022). correlation: Methods for correlation analysis [R package]. https://CRAN.R-project.org/package=correlation (Original work published 2020)
The correlation package is available on CRAN, while its latest development version is available on R-universe (from rOpenSci).
| Type | Source | Command |
|---|---|---|
| Release | CRAN | install.packages("correlation") |
| Development | R-universe | install.packages("correlation", repos = "https://easystats.r-universe.dev") |
Once you have downloaded the package, you can then load it using:
library("correlation")Tip
Instead of
library(bayestestR), uselibrary(easystats). This will make all features of the easystats-ecosystem available.To stay updated, use
easystats::install_latest().
Check out package website for documentation.
The correlation package can compute many different types of correlation, including:
✅ Pearson’s correlation
✅ Spearman’s
rank correlation
✅ Kendall’s rank
correlation
✅ Biweight midcorrelation
✅ Distance correlation
✅ Percentage bend
correlation
✅ Shepherd’s Pi
correlation
✅ Blomqvist’s coefficient
✅ Hoeffding’s D
✅ Gamma
correlation
✅ Gaussian rank
correlation
✅ Point-Biserial and biserial
correlation
✅ Winsorized correlation
✅ Polychoric correlation
✅ Tetrachoric
correlation
✅ Multilevel
correlation
An overview and description of these correlations types is available here. Moreover, many of these correlation types are available as partial or within a Bayesian framework.
The main function is correlation(),
which builds on top of cor_test()
and comes with a number of possible options.
results <- correlation(iris)
results
## # Correlation Matrix (pearson-method)
##
## Parameter1 | Parameter2 | r | 95% CI | t(148) | p
## -------------------------------------------------------------------------
## Sepal.Length | Sepal.Width | -0.12 | [-0.27, 0.04] | -1.44 | 0.152
## Sepal.Length | Petal.Length | 0.87 | [ 0.83, 0.91] | 21.65 | < .001***
## Sepal.Length | Petal.Width | 0.82 | [ 0.76, 0.86] | 17.30 | < .001***
## Sepal.Width | Petal.Length | -0.43 | [-0.55, -0.29] | -5.77 | < .001***
## Sepal.Width | Petal.Width | -0.37 | [-0.50, -0.22] | -4.79 | < .001***
## Petal.Length | Petal.Width | 0.96 | [ 0.95, 0.97] | 43.39 | < .001***
##
## p-value adjustment method: Holm (1979)
## Observations: 150The output is not a square matrix, but a (tidy) dataframe with all correlations tests per row. One can also obtain a matrix using:
summary(results)
## # Correlation Matrix (pearson-method)
##
## Parameter | Petal.Width | Petal.Length | Sepal.Width
## -------------------------------------------------------
## Sepal.Length | 0.82*** | 0.87*** | -0.12
## Sepal.Width | -0.37*** | -0.43*** |
## Petal.Length | 0.96*** | |
##
## p-value adjustment method: Holm (1979)Note that one can also obtain the full, square and redundant matrix using:
summary(results, redundant = TRUE)
## # Correlation Matrix (pearson-method)
##
## Parameter | Sepal.Length | Sepal.Width | Petal.Length | Petal.Width
## ----------------------------------------------------------------------
## Sepal.Length | | -0.12 | 0.87*** | 0.82***
## Sepal.Width | -0.12 | | -0.43*** | -0.37***
## Petal.Length | 0.87*** | -0.43*** | | 0.96***
## Petal.Width | 0.82*** | -0.37*** | 0.96*** |
##
## p-value adjustment method: Holm (1979)library(see)
results %>%
summary(redundant = TRUE) %>%
plot()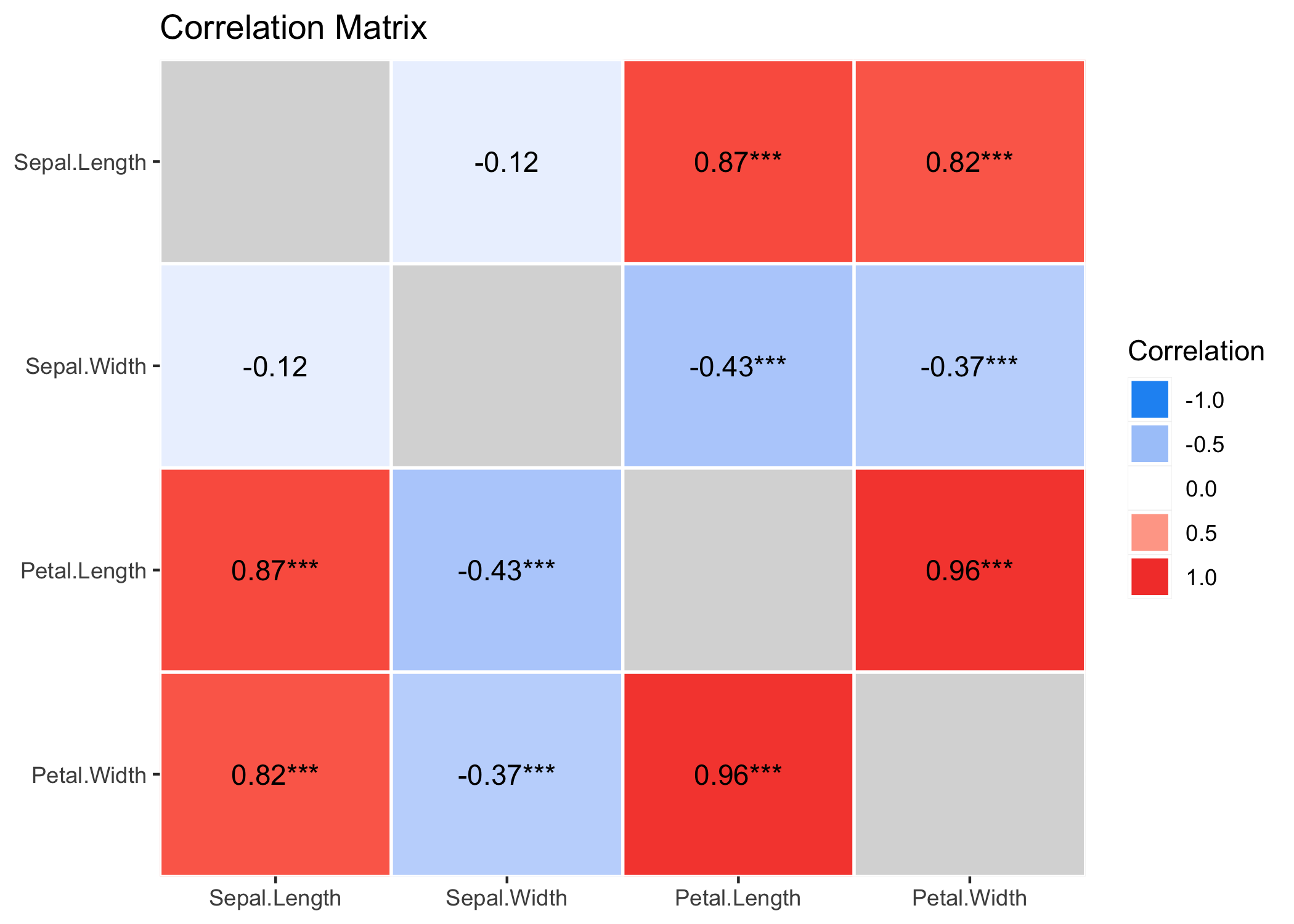
The cor_test() function, for pairwise correlations, is
also very convenient for making quick scatter plots.
plot(cor_test(iris, "Sepal.Width", "Sepal.Length"))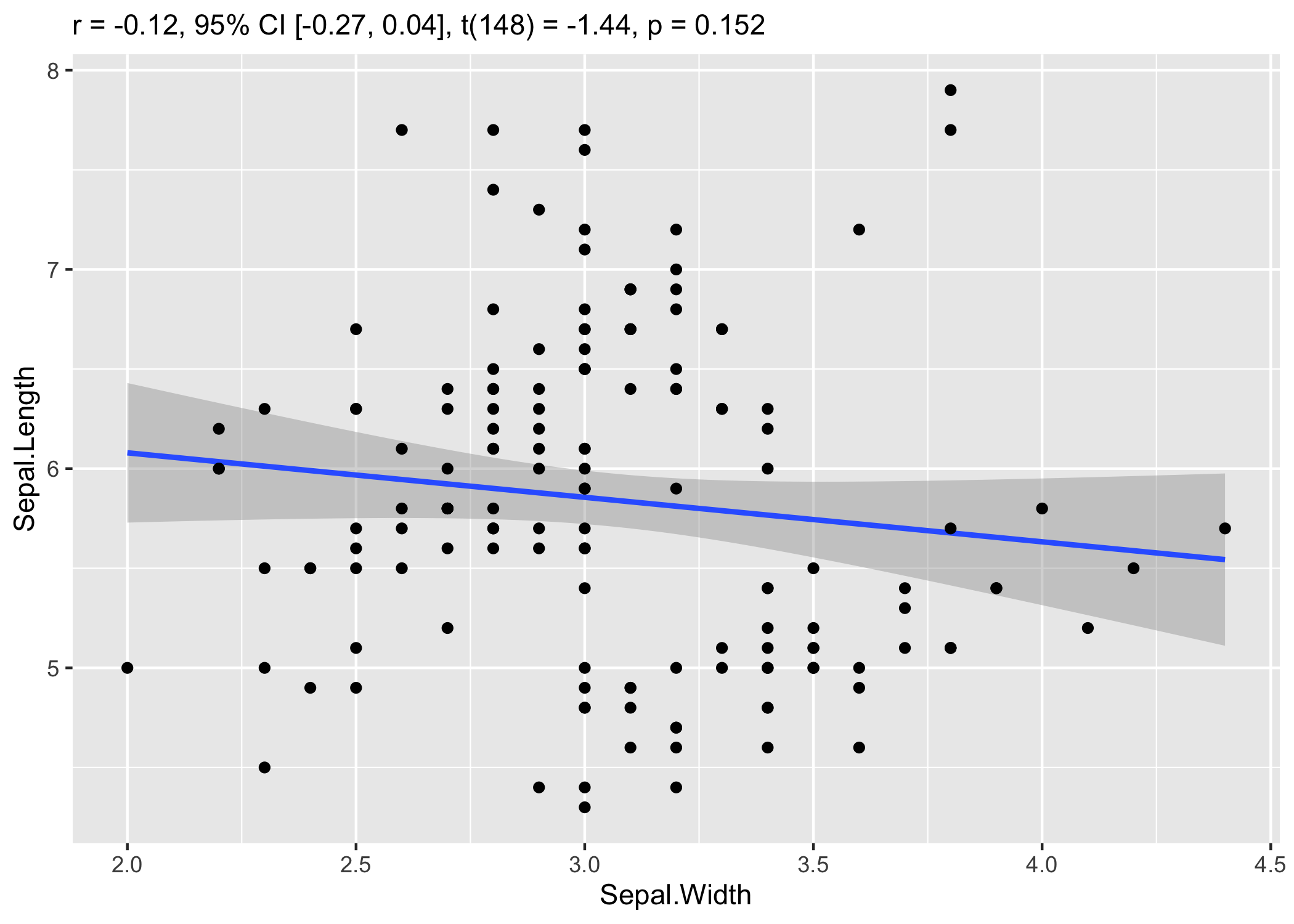
The correlation() function also supports
stratified correlations, all within the
tidyverse workflow!
iris %>%
select(Species, Sepal.Length, Sepal.Width, Petal.Width) %>%
group_by(Species) %>%
correlation()
## # Correlation Matrix (pearson-method)
##
## Group | Parameter1 | Parameter2 | r | 95% CI | t(48) | p
## ----------------------------------------------------------------------------------
## setosa | Sepal.Length | Sepal.Width | 0.74 | [ 0.59, 0.85] | 7.68 | < .001***
## setosa | Sepal.Length | Petal.Width | 0.28 | [ 0.00, 0.52] | 2.01 | 0.101
## setosa | Sepal.Width | Petal.Width | 0.23 | [-0.05, 0.48] | 1.66 | 0.104
## versicolor | Sepal.Length | Sepal.Width | 0.53 | [ 0.29, 0.70] | 4.28 | < .001***
## versicolor | Sepal.Length | Petal.Width | 0.55 | [ 0.32, 0.72] | 4.52 | < .001***
## versicolor | Sepal.Width | Petal.Width | 0.66 | [ 0.47, 0.80] | 6.15 | < .001***
## virginica | Sepal.Length | Sepal.Width | 0.46 | [ 0.20, 0.65] | 3.56 | 0.002**
## virginica | Sepal.Length | Petal.Width | 0.28 | [ 0.00, 0.52] | 2.03 | 0.048*
## virginica | Sepal.Width | Petal.Width | 0.54 | [ 0.31, 0.71] | 4.42 | < .001***
##
## p-value adjustment method: Holm (1979)
## Observations: 50It is very easy to switch to a Bayesian framework.
correlation(iris, bayesian = TRUE)
## # Correlation Matrix (pearson-method)
##
## Parameter1 | Parameter2 | rho | 95% CI | pd | % in ROPE
## --------------------------------------------------------------------------
## Sepal.Length | Sepal.Width | -0.11 | [-0.27, 0.04] | 92.70% | 42.83%
## Sepal.Length | Petal.Length | 0.86 | [ 0.82, 0.90] | 100%*** | 0%
## Sepal.Length | Petal.Width | 0.81 | [ 0.75, 0.86] | 100%*** | 0%
## Sepal.Width | Petal.Length | -0.41 | [-0.55, -0.28] | 100%*** | 0%
## Sepal.Width | Petal.Width | -0.35 | [-0.49, -0.22] | 100%*** | 0%
## Petal.Length | Petal.Width | 0.96 | [ 0.95, 0.97] | 100%*** | 0%
##
## Parameter1 | Prior | BF
## ------------------------------------------
## Sepal.Length | Beta (3 +- 3) | 0.509
## Sepal.Length | Beta (3 +- 3) | 2.14e+43***
## Sepal.Length | Beta (3 +- 3) | 2.62e+33***
## Sepal.Width | Beta (3 +- 3) | 3.49e+05***
## Sepal.Width | Beta (3 +- 3) | 5.29e+03***
## Petal.Length | Beta (3 +- 3) | 1.24e+80***
##
## Observations: 150The correlation package also supports different types of
methods, which can deal with correlations between
factors!
correlation(iris, include_factors = TRUE, method = "auto")
## # Correlation Matrix (auto-method)
##
## Parameter1 | Parameter2 | r | 95% CI | t(148) | p
## -------------------------------------------------------------------------------------
## Sepal.Length | Sepal.Width | -0.12 | [-0.27, 0.04] | -1.44 | 0.452
## Sepal.Length | Petal.Length | 0.87 | [ 0.83, 0.91] | 21.65 | < .001***
## Sepal.Length | Petal.Width | 0.82 | [ 0.76, 0.86] | 17.30 | < .001***
## Sepal.Length | Species.setosa | -0.72 | [-0.79, -0.63] | -12.53 | < .001***
## Sepal.Length | Species.versicolor | 0.08 | [-0.08, 0.24] | 0.97 | 0.452
## Sepal.Length | Species.virginica | 0.64 | [ 0.53, 0.72] | 10.08 | < .001***
## Sepal.Width | Petal.Length | -0.43 | [-0.55, -0.29] | -5.77 | < .001***
## Sepal.Width | Petal.Width | -0.37 | [-0.50, -0.22] | -4.79 | < .001***
## Sepal.Width | Species.setosa | 0.60 | [ 0.49, 0.70] | 9.20 | < .001***
## Sepal.Width | Species.versicolor | -0.47 | [-0.58, -0.33] | -6.44 | < .001***
## Sepal.Width | Species.virginica | -0.14 | [-0.29, 0.03] | -1.67 | 0.392
## Petal.Length | Petal.Width | 0.96 | [ 0.95, 0.97] | 43.39 | < .001***
## Petal.Length | Species.setosa | -0.92 | [-0.94, -0.89] | -29.13 | < .001***
## Petal.Length | Species.versicolor | 0.20 | [ 0.04, 0.35] | 2.51 | 0.066
## Petal.Length | Species.virginica | 0.72 | [ 0.63, 0.79] | 12.66 | < .001***
## Petal.Width | Species.setosa | -0.89 | [-0.92, -0.85] | -23.41 | < .001***
## Petal.Width | Species.versicolor | 0.12 | [-0.04, 0.27] | 1.44 | 0.452
## Petal.Width | Species.virginica | 0.77 | [ 0.69, 0.83] | 14.66 | < .001***
## Species.setosa | Species.versicolor | -0.88 | [-0.91, -0.84] | -22.43 | < .001***
## Species.setosa | Species.virginica | -0.88 | [-0.91, -0.84] | -22.43 | < .001***
## Species.versicolor | Species.virginica | -0.88 | [-0.91, -0.84] | -22.43 | < .001***
##
## p-value adjustment method: Holm (1979)
## Observations: 150It also supports partial correlations (as well as Bayesian partial correlations).
iris %>%
correlation(partial = TRUE) %>%
summary()
## # Correlation Matrix (pearson-method)
##
## Parameter | Petal.Width | Petal.Length | Sepal.Width
## -------------------------------------------------------
## Sepal.Length | -0.34*** | 0.72*** | 0.63***
## Sepal.Width | 0.35*** | -0.62*** |
## Petal.Length | 0.87*** | |
##
## p-value adjustment method: Holm (1979)Such partial correlations can also be represented as Gaussian Graphical Models (GGM), an increasingly popular tool in psychology. A GGM traditionally include a set of variables depicted as circles (“nodes”), and a set of lines that visualize relationships between them, which thickness represents the strength of association (see Bhushan et al., 2019).
library(see) # for plotting
library(ggraph) # needs to be loaded
plot(correlation(mtcars, partial = TRUE)) +
scale_edge_color_continuous(low = "#000004FF", high = "#FCFDBFFF")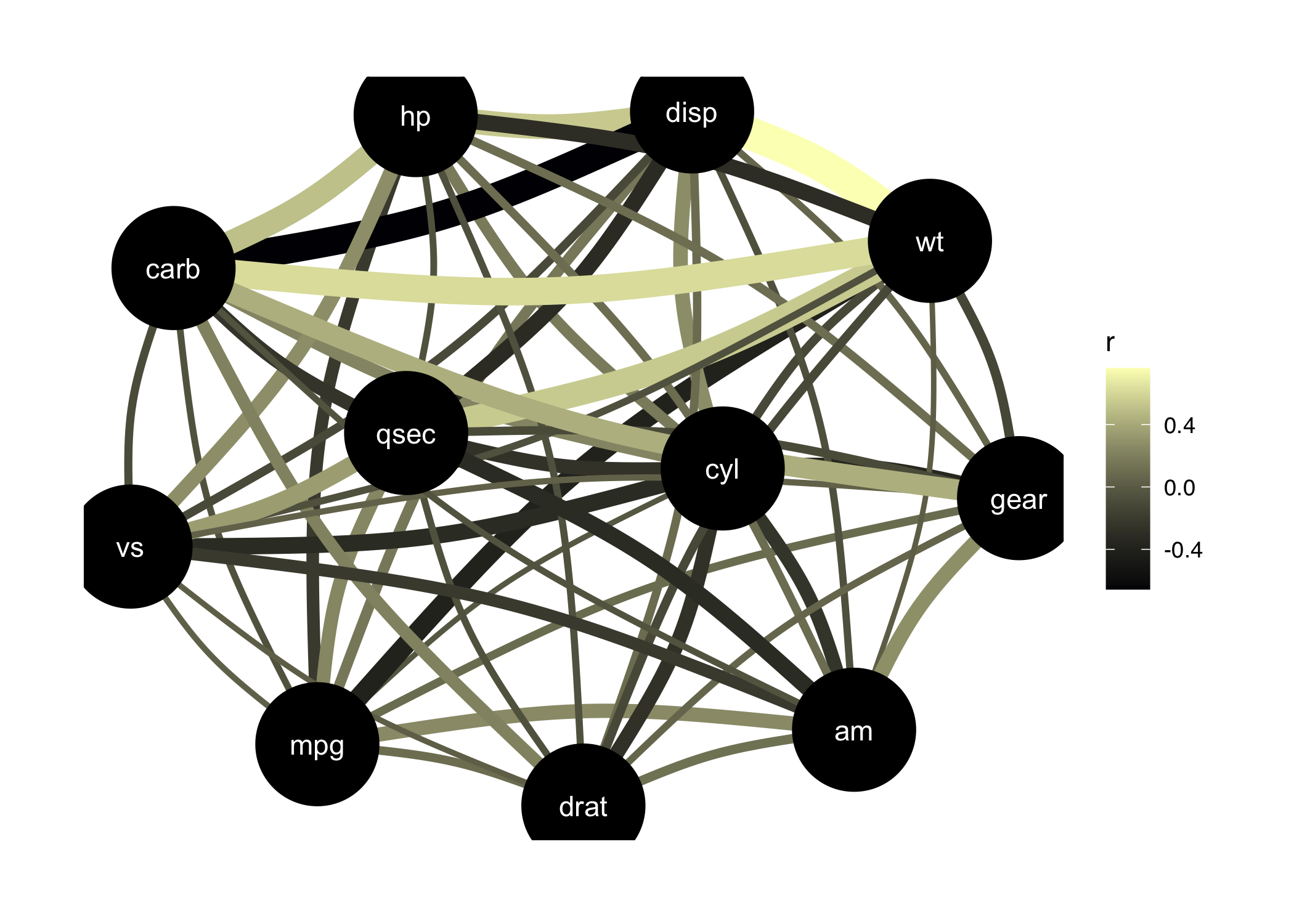
It also provide some cutting-edge methods, such as Multilevel (partial) correlations. These are are partial correlations based on linear mixed-effects models that include the factors as random effects. They can be see as correlations adjusted for some group (hierarchical) variability.
iris %>%
correlation(partial = TRUE, multilevel = TRUE) %>%
summary()
## # Correlation Matrix (pearson-method)
##
## Parameter | Petal.Width | Petal.Length | Sepal.Width
## -------------------------------------------------------
## Sepal.Length | -0.17* | 0.71*** | 0.43***
## Sepal.Width | 0.39*** | -0.18* |
## Petal.Length | 0.38*** | |
##
## p-value adjustment method: Holm (1979)However, if the partial argument is set to
FALSE, it will try to convert the partial coefficient into
regular ones.These can be converted back to full
correlations:
iris %>%
correlation(partial = FALSE, multilevel = TRUE) %>%
summary()
## Parameter | Petal.Width | Petal.Length | Sepal.Width
## -------------------------------------------------------
## Sepal.Length | 0.36*** | 0.76*** | 0.53***
## Sepal.Width | 0.47*** | 0.38*** |
## Petal.Length | 0.48*** | |In case you want to file an issue or contribute in another way to the package, please follow this guide. For questions about the functionality, you may either contact us via email or also file an issue.
Please note that this project is released with a Contributor Code of Conduct. By participating in this project you agree to abide by its terms.